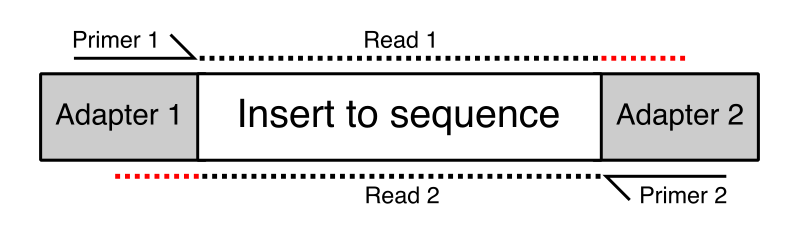
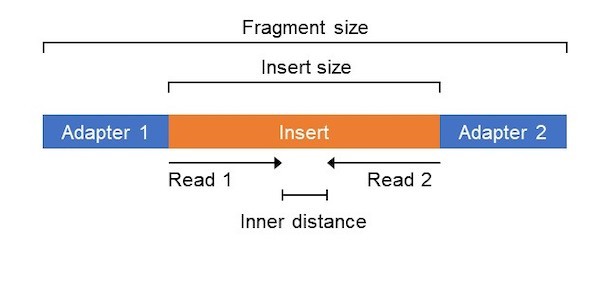
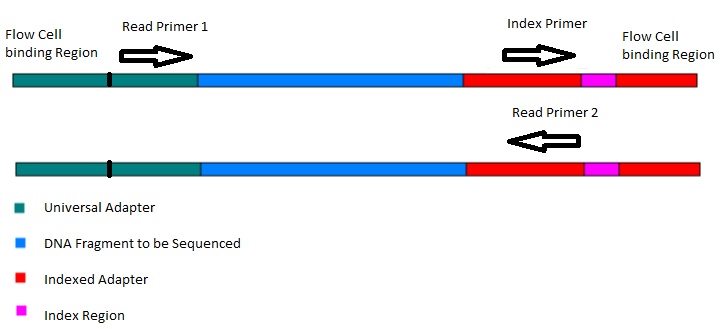
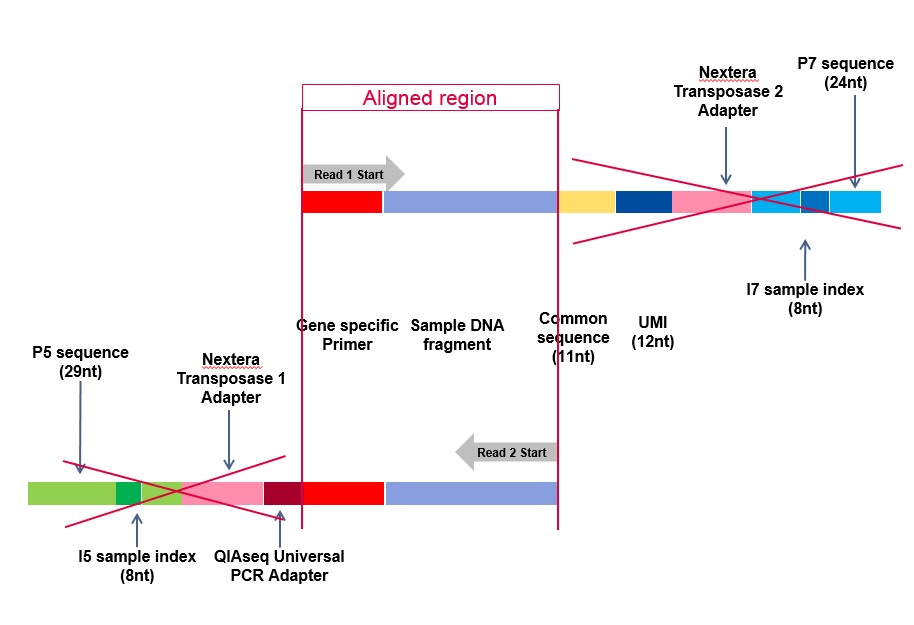
Adapter trimming: Why are adapter sequences trimmed from only the 3' ends of reads - Illumina Knowledge

Adapter trimming: Why are adapter sequences trimmed from only the 3' ends of reads - Illumina Knowledge
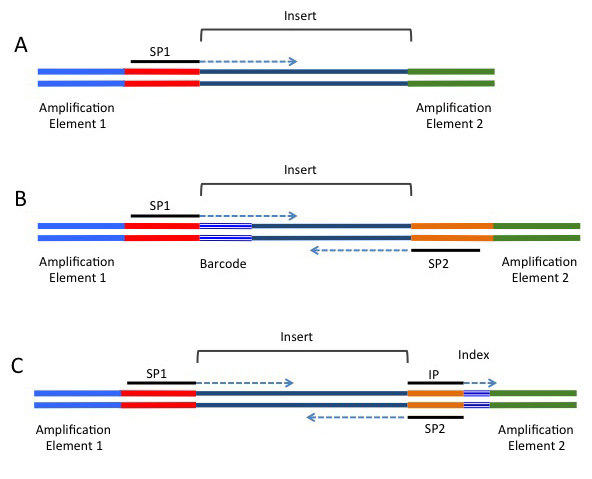
Biotech7005 | The practical material from the course Biotech 7005: Bioinformatics and Systems Modelling

Adapter trimming: Why are adapter sequences trimmed from only the 3' ends of reads - Illumina Knowledge
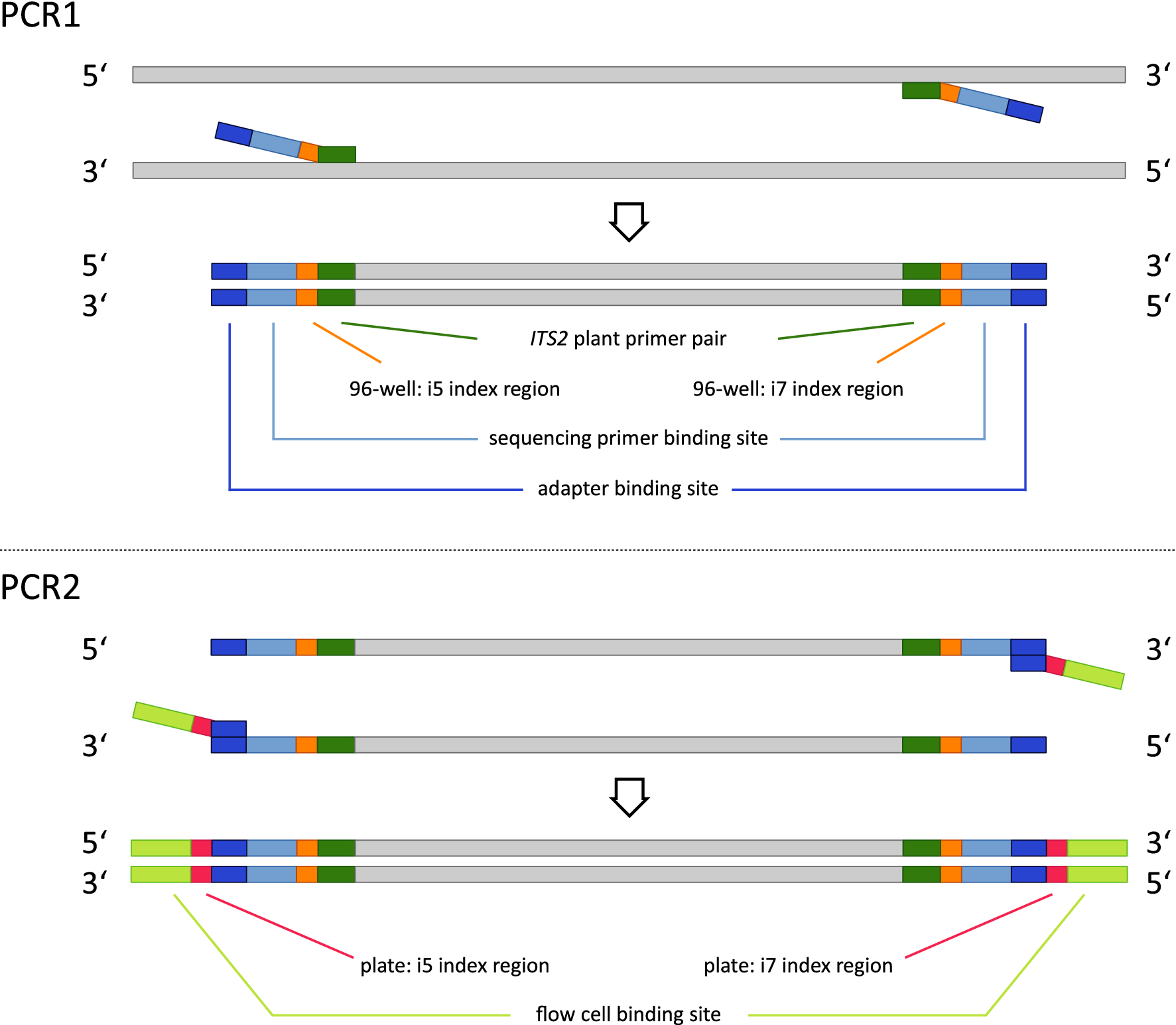
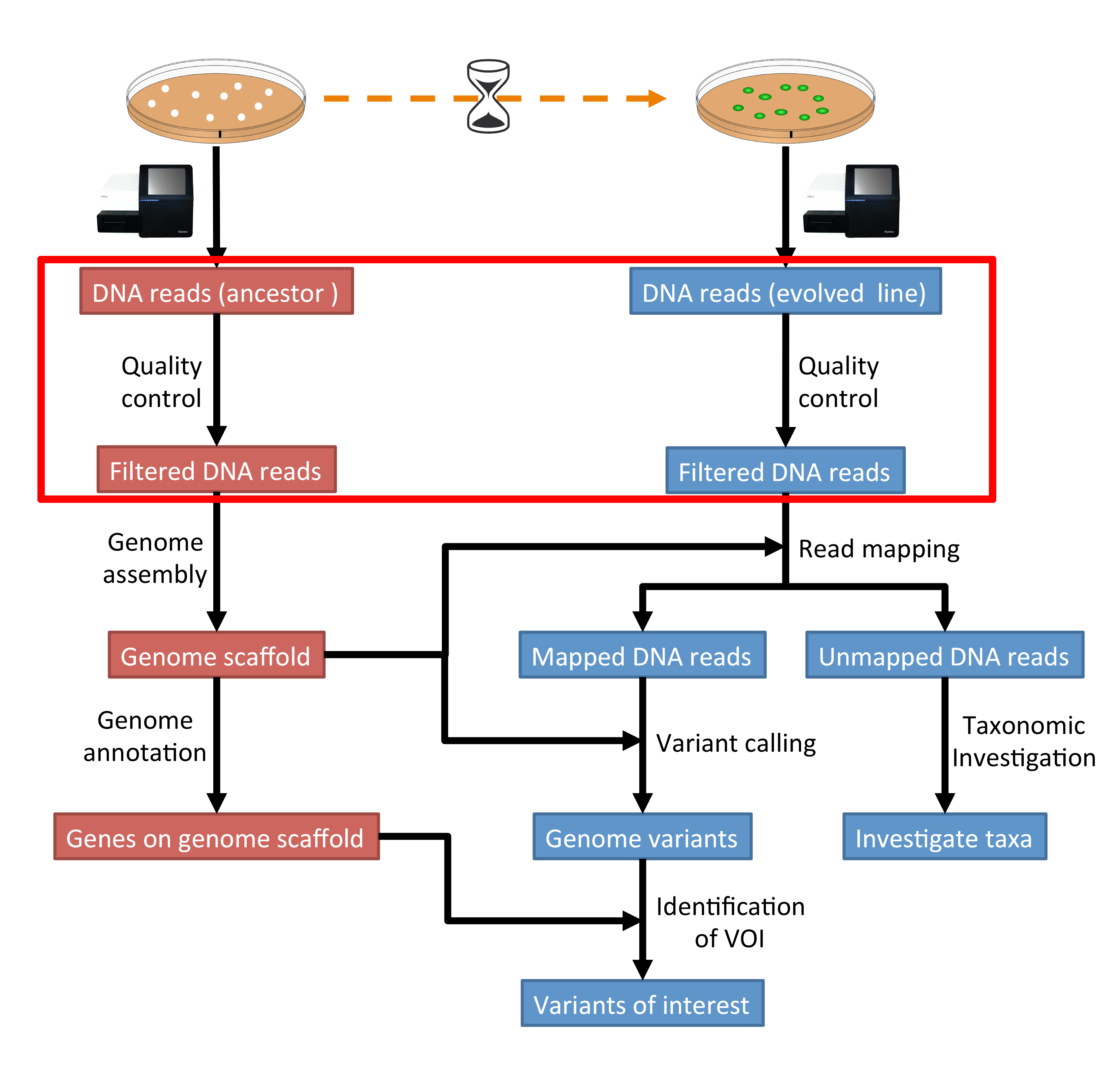
Handling of targeted amplicon sequencing data focusing on index hopping and demultiplexing using a nested metabarcoding approach in ecology | Scientific Reports

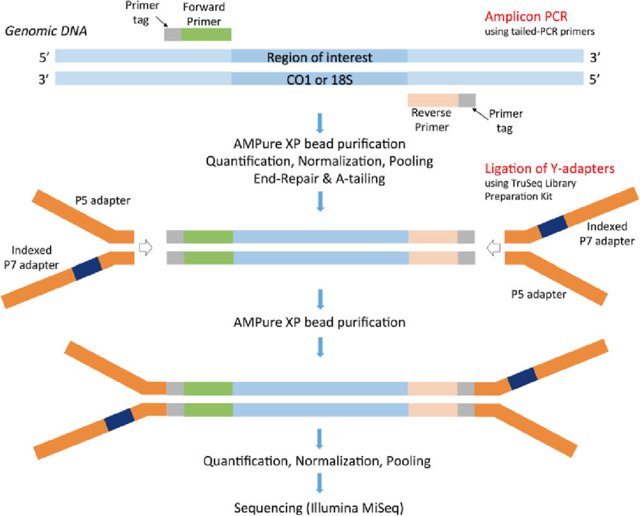
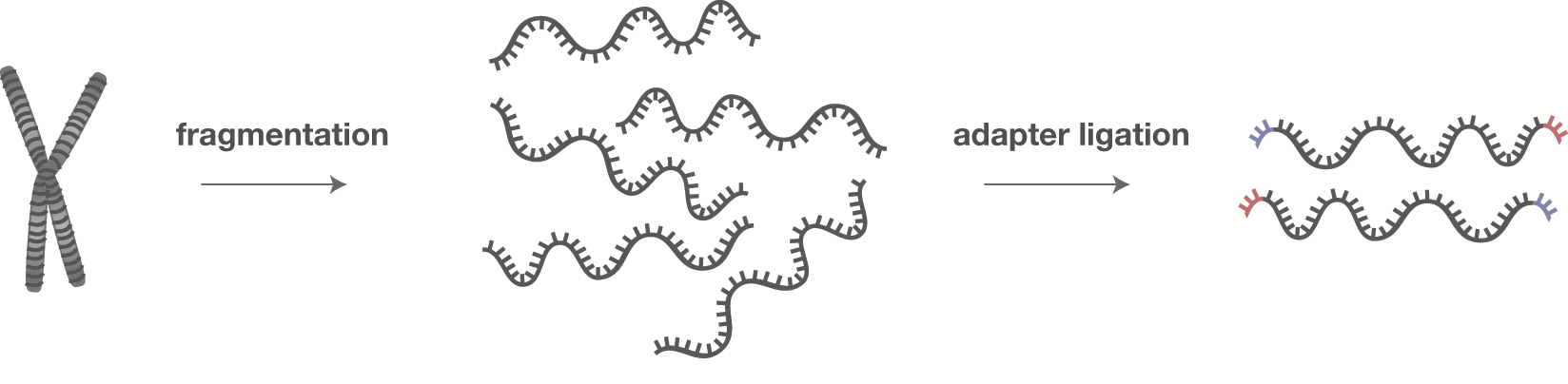
Preparation of DNA Sequencing Libraries for Illumina Systems—6 Key Steps in the Workflow | Thermo Fisher Scientific - JP