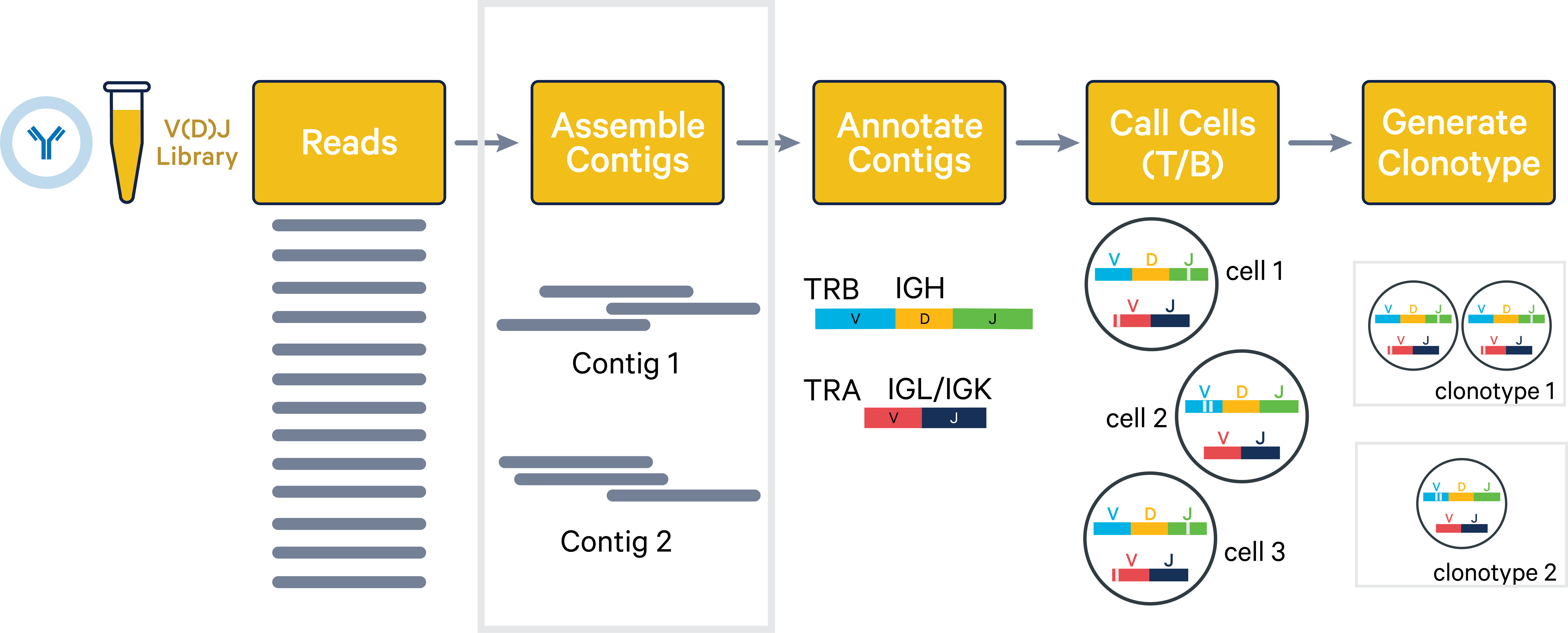
SeqPurge: highly-sensitive adapter trimming for paired-end NGS data | BMC Bioinformatics | Full Text
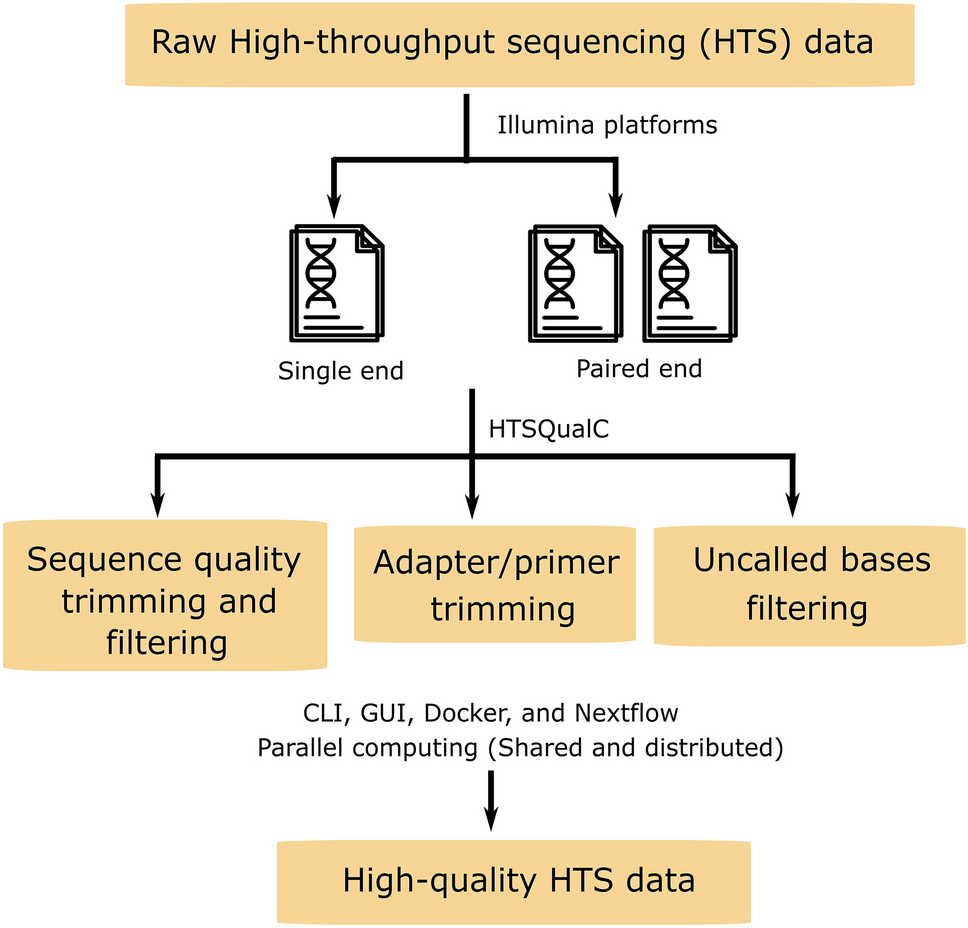
HTSQualC is a flexible and one-step quality control software for high-throughput sequencing data analysis | Scientific Reports

Biomolecules | Free Full-Text | High-Throughput Identification of Adapters in Single-Read Sequencing Data

Biology | Free Full-Text | FLEXBAR—Flexible Barcode and Adapter Processing for Next-Generation Sequencing Platforms

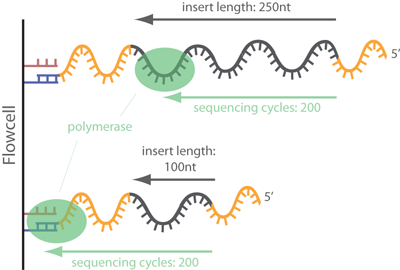
Adapter trimming: Why are adapter sequences trimmed from only the 3' ends of reads - Illumina Knowledge

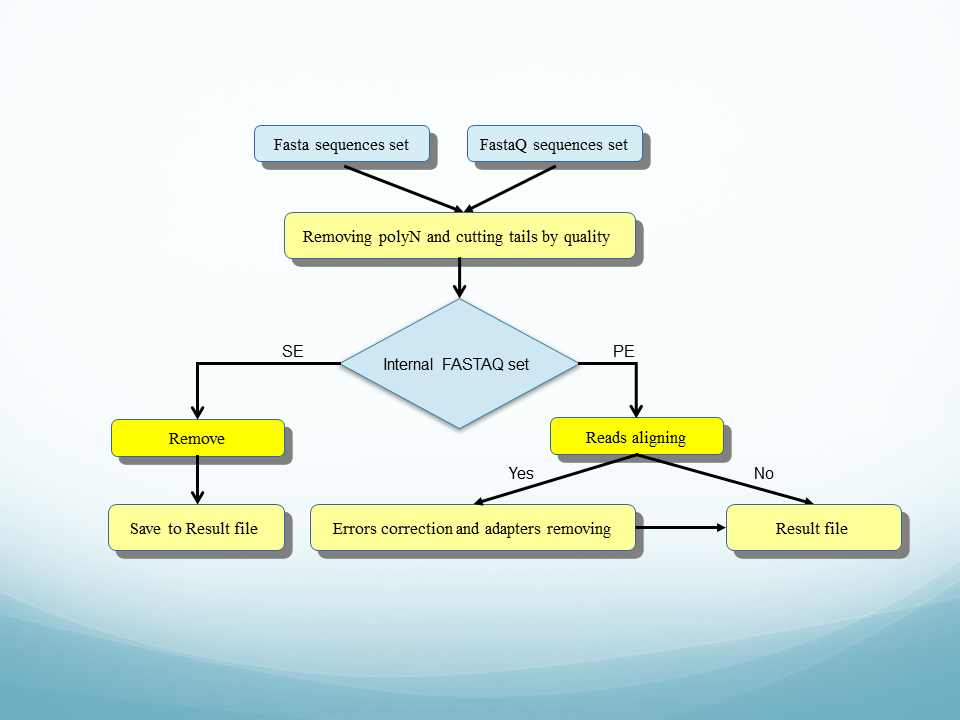
Adapter trimming for single end sRNA-seq data. The reads processed by... | Download Scientific Diagram
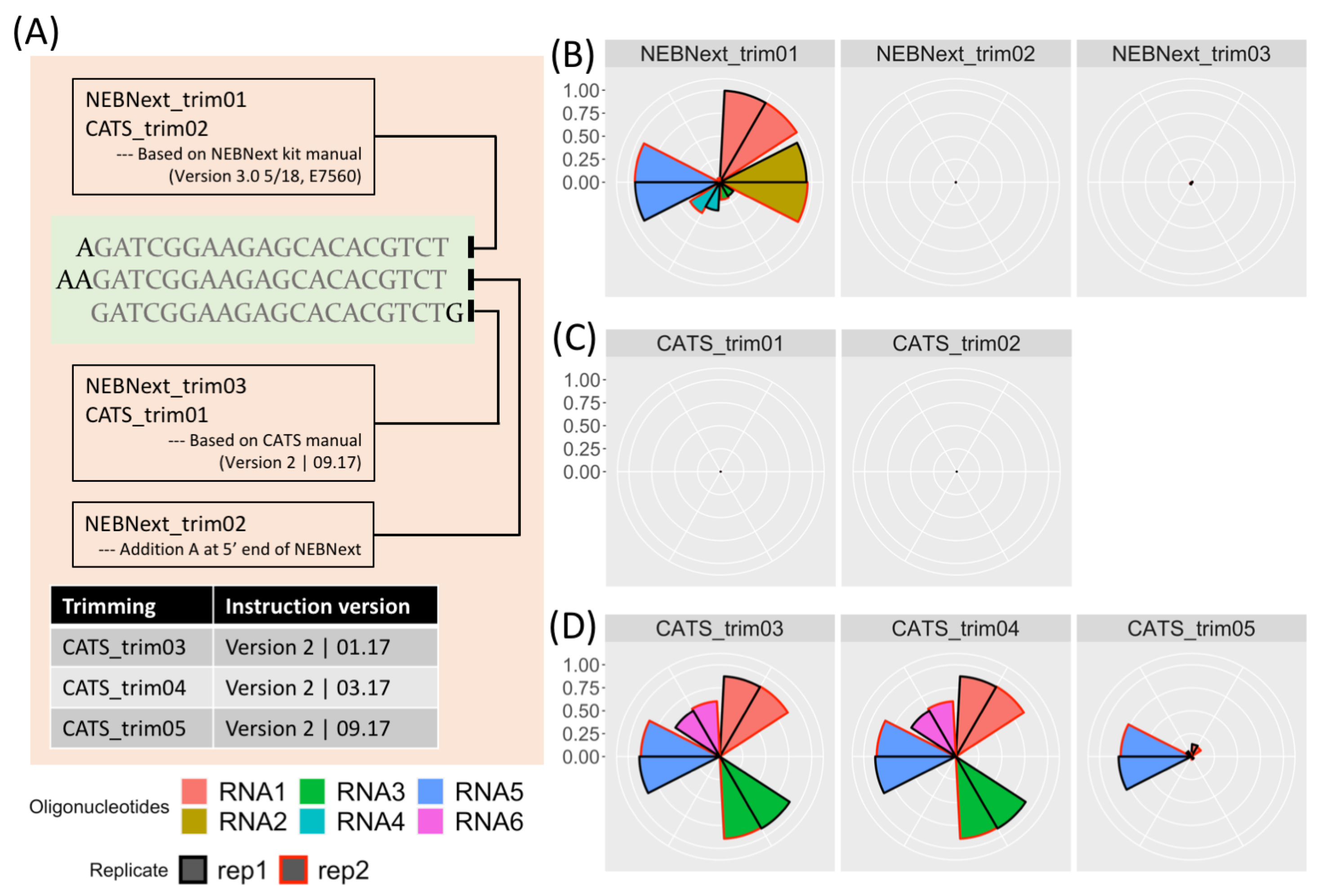
ncRNA | Free Full-Text | Accurate Adapter Information Is Crucial for Reproducibility and Reusability in Small RNA Seq Studies
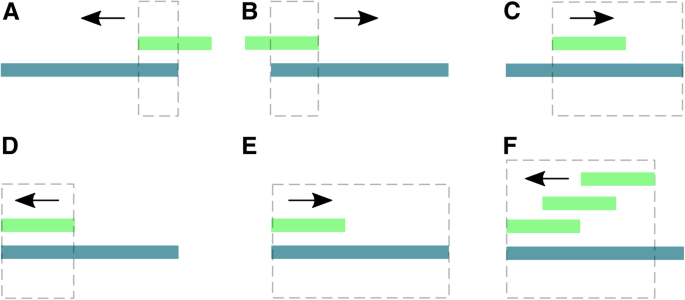




![PDF] leeHom: adaptor trimming and merging for Illumina sequencing reads | Semantic Scholar PDF] leeHom: adaptor trimming and merging for Illumina sequencing reads | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/247973613dfa3c42eb437ec9f5269202595f96af/2-Figure1-1.png)